 Vector took a moment this morning at the Boston Children’s Hospital Global Pediatric Innovation Summit + Awards to catch up with the Gene Discovery Core at the Manton Center for Orphan Disease Research. Its exhibition table doesn’t have fancy mannequins or flashy screens, but this team is rocking genetics and genomics, one patient at a time.
Vector took a moment this morning at the Boston Children’s Hospital Global Pediatric Innovation Summit + Awards to catch up with the Gene Discovery Core at the Manton Center for Orphan Disease Research. Its exhibition table doesn’t have fancy mannequins or flashy screens, but this team is rocking genetics and genomics, one patient at a time.
The usual methods for finding disease-causing genes don’t work for many patients who walk in the doors of Boston Children’s, or who mail in samples from all over the world. They may be one of just a handful of patients in the world with their condition—which may not even have a name yet. Full story »

New methods can find mutations that strike just 1 in 10 cells in a sample.
It’s become clear that our DNA is far from identical from cell to cell and that disease-causing mutations can happen in some of our cells and not others,
arising at some point after we’re conceived. These so-called somatic mutations—affecting just a percentage of cells—are subtle and easy to overlook, even with next-generation genomic sequencing. And they could be more important in neurologic and psychiatric disorders than we thought.
“There are two kinds of somatic mutations that get missed,” says Christopher Walsh, MD, PhD, chief of Genetics and Genomics at Boston Children’s Hospital. “One is mutations that are limited to specific tissues: If we do a blood test, but the mutation is only in the brain, we won’t find it. Other mutations may be in all tissues but in only a fraction of the cells—a mosaic pattern. These could be detectable through a blood test in the clinic but aren’t common enough to be easily detectable.”
That’s where deep sequencing comes in. Reporting last month in The New England Journal of Medicine, Walsh and postdoctoral fellow Saumya Jamuar, MD, used the technique in 158 patients with brain malformations of unknown genetic cause, some from Walsh’s clinic, who had symptoms such as seizures, intellectual disability and speech and language impairments. Full story »

A good biomarker is one whose levels go up or down as a patient’s disease worsens or wanes. A great biomarker also gives key insights into disease development. A really great biomarker does both of these things and also serves as a treatment target.
With a protein called netrin-1, Edward Smith, MD, and Michael Klagsbrun, PhD, seem to have hit the trifecta. In a recent paper in Cancer Research, they report a clear relationship between urine netrin levels and medulloblastoma, the most common malignant brain tumor of children.
And show that netrin fuels the tumor’s invasion into healthy brain tissue.
And that blocking netrin may, at least in the laboratory, check the tumor’s spread. Full story »

Emmie Mendes was lucky enough to be diagnosed before age 3, but many families face a much longer journey.
At first, Corrie and Adam Mendes thought their daughter Emmie had an inner ear problem. She was late with several early milestones, including walking, and when she did walk, she often lost her balance. The family pediatrician sent them to a neurologist, who ordered a brain MRI and diagnosed her with pachygyria, a rare condition in which the brain is smoother than normal, lacking its usual number of folds.
Additionally, Emmie’s ventricles, the fluid-filled cushions around the brain, looked enlarged, so the neurologist recommended brain surgery to install a shunt to drain off fluid. He advised Corrie and Adam that Emmie’s life expectancy would be greatly reduced.
As Corrie recounts on her blog, Emmie’s Story, she went online and came across the research laboratory of Christopher Walsh, MD, PhD, at Boston Children’s Hospital. The lab does research on brain malformations and has an affiliated Brain Development and Genetics Clinic that can provide medical care.
After Walsh’s team reviewed Emmie’s MRI scan, genetic counselor Brenda Barry invited the family up from Florida. Full story »
A randomized trial will soon test whether web-based updates from parents and teachers improve outcomes in ADHD, autism and more.
Eugenia Chan, MD, MPH, is a developmental-behavioral pediatrician and health services researcher in the Division of Developmental Medicine at Boston Children’s Hospital. She runs the Developmental Medicine Center’s ADHD Program and is co-developer of ICISS Health, a web-based disease monitoring and management system.
When I set out with my collaborator Eric Fleegler, MD, MPH, to build a web-based tracking system for children with attention deficit hyperactivity disorder (ADHD), we focused on a single problem—getting parents and teachers to fill out symptom questionnaires in time to help doctors make informed clinical decisions at follow-up visits. We had no inkling of the possibilities that this kind of software platform could hold, or how it might grow in the future. Full story »
 What happens when you try to scale up a successful quality initiative? Eric Fleegler, MD, MPH, and Eugenia Chan, MD, MPH, are facing that challenge with ICISS, their web-based system that went quickly from ideation to adoption by 3,000 patients with attention deficit hyperactivity disorder (ADHD) and their families.
What happens when you try to scale up a successful quality initiative? Eric Fleegler, MD, MPH, and Eugenia Chan, MD, MPH, are facing that challenge with ICISS, their web-based system that went quickly from ideation to adoption by 3,000 patients with attention deficit hyperactivity disorder (ADHD) and their families.
ICISS enables parents, teachers and patients to give online updates on medications, symptoms and school performance in close to real time, then packages that data for clinicians in a visual, actionable fashion. But tasked with introducing ICISS into four other clinics at Boston Children’s Hospital—autism, asthma, depression and epilepsy—a raft of practical, legal and philosophical questions came up about how to handle these patient-generated health data. For example:
- How should we inform families that they need to contact their provider directly with immediate concerns?
- What if a parent indicates that a child is at risk of self-harm, and how can we manage this in a timely manner?
- How can clinics afford to hire additional staff to screen and manage alerts from ICISS when this activity is non-reimbursable?
- What is the obligation of the provider if actionable data show up months in advance of the scheduled visit?
Fleegler and Chan discuss the challenges and lessons learned in our sister publication, Innovation Insider. We’d be interested to hear from others facing similar questions in handling patient-generated health data.
 Last November, the U.S. Food and Drug Administration issued a “cease and desist” order to 23andMe, a major purveyor of direct-to-consumer (DTC) genetic testing. In its letter to the company—issued after three prior warnings—the FDA reiterated its view that 23andMe’s Personal Genome Service (PGS) constitutes a medical device requiring further premarket evaluation:
Last November, the U.S. Food and Drug Administration issued a “cease and desist” order to 23andMe, a major purveyor of direct-to-consumer (DTC) genetic testing. In its letter to the company—issued after three prior warnings—the FDA reiterated its view that 23andMe’s Personal Genome Service (PGS) constitutes a medical device requiring further premarket evaluation:
FDA is concerned about the public health consequences of inaccurate results from the PGS device…we still do not have any assurance that the firm has analytically or clinically validated the PGS for its intended uses.
The FDA’s order, based on potential rather than actual medical harm, has generated a great deal of controversy. In a recent critique published in Nature, Robert Green, MD, MPH, of the Partners HealthCare Center for Personalized Genetic Medicine, and Nita Farahany, PhD, JD, of the Duke Institute for Genome Sciences and Policy, argued against regulating DTC genomic interpretation services as medical devices:
… doing so could put FDA regulations in greater tension with the First Amendment of the US Constitution, which protects the rights of individuals to receive information, and of ‘commercial speech’ ….the agency should avoid restricting consumer genomic testing unless faced with empirical evidence of harm. Full story »
 In 2012, Boston Children’s Hospital held the international CLARITY Challenge—an invitation to interpret genomic sequence data from three children with rare diseases and provide a meaningful, actionable report for clinicians and families. (Click for more background on the children, findings and winners.)
In 2012, Boston Children’s Hospital held the international CLARITY Challenge—an invitation to interpret genomic sequence data from three children with rare diseases and provide a meaningful, actionable report for clinicians and families. (Click for more background on the children, findings and winners.)
The full proceedings, published March 25 in Genome Biology, concluded that while the technical approaches were markedly similar from center to center, the costs, efficiency and scalability were not. Most variable, and most in need of future work, was the quality of the clinical reporting and patient consenting process. The exercise also underscored the need for medical expertise to bring meaning to the genomic data.
That was CLARITY 1. CLARITY 2, focusing on cancer genomics in children, promises to be exponentially more complex. Full story »
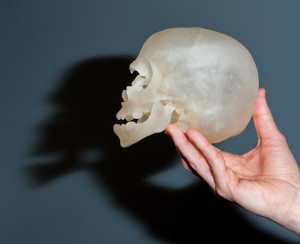 A picture may be worth a thousand words, but there’s something about holding an object in your hands that’s worth so much more. I realized this when John Meara, MD, DMD, handed me the skull of one of his patients.
A picture may be worth a thousand words, but there’s something about holding an object in your hands that’s worth so much more. I realized this when John Meara, MD, DMD, handed me the skull of one of his patients.
I turned it over in my hands while Meara, Boston Children’s Hospital’s plastic surgeon-in-chief, pointed out features like the cranium’s asymmetric shape and the face’s malformed left orbit.
Mind you, it wasn’t actually Meara’s patient’s skull in my hands. In reality, I was holding a high-resolution, plastic 3D model printed from the patient’s CT scans.
The printer that made that model—and several other models I saw in the last month—is the centerpiece of a new in-house 3D printing service being built by Peter Weinstock, MD, PhD, and Boston Children’s Simulator Program.
3D printing technology has exploded in the last few years, to the point where anyone can buy a 3D printer like the MakerBot for a couple of thousand dollars or order 3D printed products from services like Shapeways. Adobe even recently added 3D printing support to Photoshop.
And 3D printing is already making a mark on medicine. Full story »

Three clinics are pre-testing children for genetic variants likely to affect their response to drugs.
In 2009,
The New England Journal of Medicine reported the
case of an otherwise healthy 2-year-old boy in Canada who died after surgery. He had received a codeine dose in the recommended range, but an autopsy revealed that morphine (a product of codeine metabolism) had built up to toxic levels in his blood and likely depressed his breathing. Genetic profiling revealed him to be an “ultrarapid codeine metabolizer,” due to a genetic variation in an enzyme known as CYP2D6, part of the cytochrome P-450 family.
While codeine is no longer used at Boston Children’s Hospital, it’s this kind of genetic profiling that Shannon Manzi, PharmD, would someday like to offer to all patients—before a drug is prescribed.
Not all people respond the same way to drugs. The results of randomized clinical trials—considered the gold standard for drug testing—often produce a dose range that worked for the majority of the patients in the study. They don’t take people’s individuality into account, and that individuality can dramatically affect drug efficacy and toxicity.
Adverse reactions are more common than you might think. Full story »
 Vector took a moment this morning at the Boston Children’s Hospital Global Pediatric Innovation Summit + Awards to catch up with the Gene Discovery Core at the Manton Center for Orphan Disease Research. Its exhibition table doesn’t have fancy mannequins or flashy screens, but this team is rocking genetics and genomics, one patient at a time.
Vector took a moment this morning at the Boston Children’s Hospital Global Pediatric Innovation Summit + Awards to catch up with the Gene Discovery Core at the Manton Center for Orphan Disease Research. Its exhibition table doesn’t have fancy mannequins or flashy screens, but this team is rocking genetics and genomics, one patient at a time.














