With the launch this summer of Emulate Inc., organs-on-chips—a disease-modeling platform we’ve covered several times on Vector—made the jump from academic to commercial development.
Though developed at the Wyss Institute for Biologically Inspired Engineering, the chips’ story actually began more than 20 years ago in Boston Children’s Hospital’s Vascular Biology Program (VBP). It’s a story that brings together characters from multiple fields and emerges from one fundamental concept: that mechanical forces are critical to the function and fate of cells, tissues and organs.
Force
When he was a doctoral student, Donald Ingber, MD, PhD—founding director of the Wyss Institute, Judah Folkman Professor of Vascular Biology at Harvard Medical School and a senior researcher in the VBP—worked to bring the idea of tensegrity (roughly speaking, that compression and tension work within a structure to help it maintain its shape and integrity) into cellular biology.
Ingber predicted that tensegrity enables cells to sense mechanical forces through specific surface proteins that link the cell’s external environment (especially the extracellular matrix) to its internal cytoskeleton. (In a 1993 paper in Science, he showed that surface proteins called integrins constitute that link.)
But once in his Boston Children’s laboratory, Ingber realized he had to find a way to sell the concept in an era when molecular growth factors were considered the last word in cell function and tissue development.
So he turned to Harvard chemistry professor George Whitesides, PhD, who was working on inexpensive methods for manufacturing microchips.
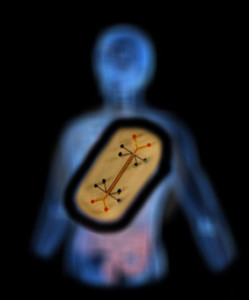
Gut-on-a-chip, with cells forming structures mimicking those of the natural intestines. (Credit: Wyss Institute)
“His team was etching patterns onto a chip and pouring on a silicon-rubber polymer,” Ingber explains. “When you peeled it off, you had a rubber mold that retained the surface topography of the chip down to the nanometer scale, the same scale at which living cells and tissues work.”
Whitesides used the molds like ink stamps for rapid and reproducible computer chip manufacturing, inking them with chemicals that he could use to stamp out the same circuit pattern over and over. Ingber, on the other hand, inked the molds with proteins and stamped out little cell-sized islands of extracellular matrix proteins of different shapes and geographies.
That work led to Science papers in 1994 and 1997 showing that cells grown on differently shaped islands will behave differently—growing when spread flat on big islands and turning on a suicide program when crowded on tiny islands—even when cultured with the same growth factors under the same conditions. It proved that mechanical forces hold as much sway as other external influences over cell behaviors, including growth, differentiation, programmed cell death and migration.
Flow
The next breakthrough was to bring in microfluidics—the means to manipulate miniscule amounts of fluid within an experimental system. Again, Ingber collaborated with Whitesides’ lab.
“George’s group had etched ridges onto silicon, poured the polymer on it and put the resulting stamp onto a glass slide. That gave them a hollow channel,” he says. From there, it was a relatively small step to generate multiple microscopic fluid channels on a single chip.
While Whitesides and others used the technology to develop miniaturized instruments, Ingber’s lab started putting cells into the chips. By controlling fluid flow, they realized, you could control a cell’s microenvironment. “You could place cells in different positions relative to each other and deliver different chemicals to different points on the surface,” Ingber says.
That capability opened the door to manipulating cells in microenvironments similar to those in the body.
Function
The last step to true organs on chips was to apply a physiology-centered point of view. In 2007, Shuichi Takayama, PhD, at the University of Michigan, whom Ingber and Whitesides had co-mentored as a postdoctoral fellow, built what was called the very first lung-on-a-chip. Essentially, it was a microfluidic chip with channels the size of small lung airways, through which he flowed liquid droplets that mimicked mucus.
“The sound that came out of this little rubber chip was exactly the same sound I was taught in medical school to listen for with the stethoscope when you suspected a patient might have pneumonia or ‘fluid-on-the-lungs’,” Ingber recalls. “It was mind-blowing.”
A graduate student from Takayama’s group who had worked on that chip, Dongeun Huh, PhD, joined Ingber’s Boston Children’s lab as a fellow. Ingber used the opportunity to expand on Takayama’s work.
By adding different types of lung cells, Ingber’s team was able to build a living tissue-tissue interface mimicking that of the lung’s smallest air sacs, where capillaries and lung tissue come in contact.
“What makes an organ an organ isn’t just tissue,” Ingber explains. “It’s having two or more different tissues that come together and form an interface, where new functions emerge.”
And to fully capture the lung’s biological and mechanical environment, the team added simulated breathing motions, just like those of a living lung.
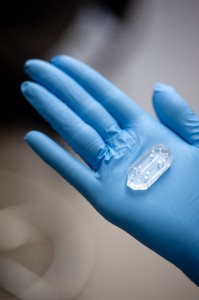
With that final ingredient, Ingber’s team found they could functionally mimic a human organ—all within a piece of clear rubber polymer the size of a matchbox.
“We get organ-level function in the chips because we put tissue/tissue interfaces, fluid flow and mechanical forces—breathing motions for the lung, peristaltic motions for the gut—together,” Ingber states. “Without all that, you don’t get in vivo physiology.”
Fruition
When the Wyss Institute first opened its doors in 2009, Ingber brought his nascent chip technology with him. In that fertile environment of biologists, engineers, fabrication specialists, machinists, commercialization experts and product developers, the chips took off.
In 2010, Ingber, Huh (who is now at the University of Pennsylvania) and their team showed that their lung-on-a-chip could simulate the lung’s response to infection and airborne particulates. In 2012, they showed that the chips could mimic a complex human disease state, pulmonary edema, and even demonstrated the ability of a new drug to prevent this life-threatening condition.
Additional chips quickly followed: a gut chip, a kidney chip, a bone marrow chip and more. Funding from the NIH, FDA, DARPA and pharmaceutical companies has fueled further development and refinement of chips, as well as automated instruments for plugging several chips together, creating human bodies-on-chips for comprehensive drug testing.
The bone marrow chip. A plug of engineered bone marrow fits into the circular section in the middle. (Credit: Wyss Institute)
While Ingber moved his engineering team to the Wyss five years ago, his cell biologists, molecular geneticists, biochemists and clinicians are all still based in his Boston Children’s lab. The “split” has, paradoxically, allowed greater collaboration: The Boston Children’s group keeps Ingber’s group grounded in basic science and clinical problems, while the Wyss-based engineers bring the technological, manufacturing and commercialization know-how.
That synergistic spirit has allowed the chips to flourish and fostered the creation of Emulate. And it infuses the Wyss Institute as a whole.
“We provide our members with access to expertise and facilities,” Ingber says. “We act like a company, but we still have the freedom of academia in that we’re not locked into any one thing. I think that increases our chances for success.”
For more success stories, join us at the Global Pediatric Innovation Summit + Awards 2014 on October 30-31 in Boston. Seats are limited, so register today at www.takingontomorrow.org. Please use the code VECTOR at checkout for a 10% discount.